This will delete the page "News". Please be certain.
September 2022: GIN for and at the Bernstein Conference
After two years of online format, the Bernstein Conference will be back to an in-person event again this year - however with a large online component to which the G-Node is contributing. As in previous years, a customized version of GIN will host all scientific posters for this conference. Moreover, this time the platform will also make all conference-related information online available to the conference attendants.
Furthermore, we will be at the conference - together with other initiatives supporting research data management at a joint RDM info booth - and present the full suite of G-Node tools and services with the latest exciting features. Drop by booth # 2, or visit our poster # 42 in poster session III.
September 2020: GIN for and at the Online Bernstein Conference
We are happy to announce, that the G-Node will be at this year's Bernstein Conference, 29. September - 01 October.
The conference will be hosted entirely online for the first time. And not only will GIN be presented online at this opportunity, a customized version of GIN will also host all scientific posters for this conference! If you want to see in action be sure to register for the conference.
The G-Node will further present its full suite of current tools and services and its latest exciting features.
You can sign up for the conference at the Bernstein conference registration site.
Check back here for details about our upcoming poster presentations and online booth hours where we hope to answer all your questions in virtual person!
June 2020: GIN updates
In the last few months we have been working hard on some redesign of the service infrastructure. We have also made some small design changes and added some new features to better serve your data hosting and publishing needs.
Anonymous access of public annexed data
It is now possible to clone a public repository and retrieve all annexed (large file) content using third party tools without a GIN account. We recommend using DataLad, a data management command line tool built on Git and Git-annex (much like the GIN client). Instructions for using DataLad with GIN and especially for cloning public repositories anonymously can be found in the GIN chapter of the DataLad Handbook.
We thank the DataLad team for their help in making this possible.
Repository page changes
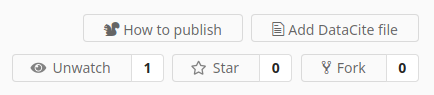
We have made some small changes to the repository header buttons (see images below):
- Users can now Star a repository as a way to show appreciation or support for a dataset. The number of people who have starred a repository is always visible in repository lists and at the top of the repository page. A list of users that have starred a repository is also visible for anyone who has access to the repository. We will soon be adding a page where users can see a list of all the repositories they have starred.
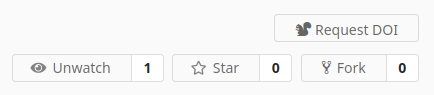
- The publication (DOI) related buttons have been changed for better clarity. The Obtaining a DOI help page has been updated accordingly.
Publishing a new dataset version
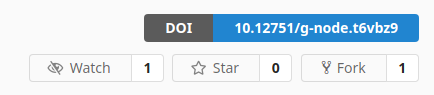
In addition to the button changes described above, we have also added the ability to request a DOI for a new version of a previously published dataset. Repositories of published datasets can be registered again by clicking the "Request DOI" button next to the DOI badge, which appears when changes to the repository are detected after initial publication.
Note that a new DOI is not required when the metadata of the published dataset have changed (e.g., for the addition of a reference).
Repositories that have been published more than once will only show the most recent DOI in the header. In the future we will display all relevant DOIs for a dataset.
Published Datasets
The pages of published datasets, found listed at https://doi.gin.g-node.org, have been redesigned. The new pages should make it easier to find links to the source repositories and published archives. We have also made it possible to browse the published datasets by keyword, which should make it easier to find datasets for specific topics.
We are also in the process of submitting the datasets to the Google Dataset Search index. The datasets should be available in the search results within the next few days.
October 2019: SfN 2019, Chicago, IL
The G-Node will once again be at this year's SfN Conference, 19-23 October, in Chicago, IL.
Find us throughout the conference at the Neuroscience in Germany Booth (2063) for demos and discussions on GIN, its new features and associated services, as well as other G-Node software and tools.
We will also be presenting demos of NIX at the INCF booth (2117).
See the schedule below for a full list of demo and poster times.
Poster
- Session: 252 (Presentation number: 252.22)
- Poster board number: BB52
- Time: Sunday, 20 October, 13:00-17:00 (Presentation time: 14:00-15:00)
GIN presentation times
- Location: Neuroscience in Germany booth (2063)
- Sessions:
- Monday, 21 October, 10:00-12:00
- Tuesday, 22 October, 15:00-17:00
- Wednesday, 23 October, 10:00-12:00 & 15:00-17:00
NIX presentation times
- Location: INCF booth (2117)
- Sessions:
- Monday, 21 October, 15:15-17:00
- Tuesday, 22 October, 09:30-11:15
See you in Chicago!
September 2019: Bernstein conference and usage stats update
GIN demos at the Bernstein Conference in Berlin
The G-Node will be attending the Bernstein Conference, which is taking place in Berlin from 18 to 20 September. Find us at our booth in the main hall. We will be providing demonstrations of GIN features, as well as new and upcoming services.
We will be available for demos, technical help, or just a friendly chat during the entire duration of the conference.
- Wednesday, 18.09, 15:00 - 18:00
- Thursday, 19.09, 09:00 - 18:00
- Friday, 20.09, 09:00 - 12:00
You can also find us at our poster (W57) during the Wednesday poster session, 16:00 - 19:00.
Usage stats update
A quick update on our usage statistics since the last time we shared (5 June):
- Total users: 402 (up from 395)
- Repositories:
- 593 total (up from 547)
- 257 public (up from 222)
- 483 primary repositories (non-forks)
- 166 public primary repositories
- Data size (all repositories): 6.1 TiB (up from 5.4 TiB)
- DOI registrations: 67 (up from 59)
- Total published data size (DOI registrations): 480 GiB (up from 455 GiB)
June 2019: Windows GUI client, Upgrades, and stats!
It's been a while since we had a news and GIN status update. Rest assured, we have all been busy working to bring you the best service we can. Here are some important new and upcoming features.
Windows GUI client
We are very proud to announce the initial release of WinGIN, a graphical user interface for managing GIN repositories and data on Windows. You can find download and installation instructions on the WinGIN page.
WinGIN was developed by Fabian Woltermann and Jiří Vaněk and we're very happy to share the product of their efforts with you. Of course the journey doesn't end here. New features and improvements will continue to come as we add new capabilities to our services and consider new ways to help you all manage your research data.
Upgrades (and downtime)
We've been preparing some updates to the GIN services that will improve the user experience for everyone. These include both software and hardware (and storage!) upgrades. For the roll-out we will need to briefly interrupt the service availability. This will occur on Monday, 24 June, 2019 at 15:00 CEST (13:00 UTC) and should not last more than a few hours. We expect all services to be back up and running by 18:00 CEST (16:00 UTC) on the same day.
Main service downtime:
- Monday, 24 June, 2019 13:00 UTC (15:00 CEST) until 16:00 UTC (18:00 CEST)
- Access to DOI registrations will not be affected
If this schedule severely impacts your work, please contact us at gin@g-node.org
Usage stats
We're happy to share usage statistics for GIN and we intend to keep you up-to-date with more frequent and detailed updates about the state of GIN going forward. Some general stats to kick off the tradition are listed below:
- Total users: 395
- Repositories: 547 (222 public)
- Data size (all repositories): 5.4 TiB
- DOI registrations: 59
- Total published data size (DOI registrations): 455 GiB
NWG 2019, Goettingen
An upcoming face to face with the G-Node, presenting new GIN features and other exciting G-Node projects at this years NWG, Goettingen Meeting of the German Neuroscience Society, March 20-23 2019.
More details about upcoming G-Node events at the conference can be found here once they are available.
SfN 2018, San Diego
The G-Node will be at this years SfN Conference, Nov 03 - Nov 07, in San Diego!
Find us throughout the conference at the Bernstein Booth for thrilling GIN demos and exciting discussions how GIN can help you managing your scientific data!
You can find us at the booths of the INCF and the Bernstein Network:
Booth 4417 (INCF)
- Tuesday, November 06, 15:00-17:00
Booth 4325 (Neuroscience in Germany)
- Sunday, November 04, 10:00-12:00 and 15:00-17:00
- Monday, November 05, 10:00-12:00 and 15:00-17:00
- Wednesday, November 07, 10:00-12:00 and 15:00-17:00
You can also casually walk by our Poster
"Achieving reproducible data workflows: Lightweight tools for safe and efficient data management" 527.07 / Board MMM24 Tuesday, November 06, 08:00-12:00
See you in San Diego!
Bernstein Conference 2018
You can still find the G-Node tech team for demos and any questions you might have for us at the Bernstein Conference 2018 in beautiful Berlin! You can visit us at any time at the Booth of the Bernstein Network.
- Wednesday, 26.09, 09:00 - 18:00
- Thursday, 27.09, 09:00 - 18:00
- Friday, 28.09, 09:00 - 13:00
Looking forward to help you get GIN!
FENS 2018

Find G-Node tech demos@FENSForum 2018 both at the booths of the INCF and the Bernstein Network. See detailed schedule below:
Booth 15 (INCF)
- Monday, July 9, 14:00-15:30
- Tuesday, July 10, 09:30-11:00
Booth 36 (Bernstein Network)
- Sunday, July 8, 14:00-16:00
- Monday, July 9, 11:00-13:00
- Tuesday, July 10, 14:00-16:00
- Wednesday, July 11, 11:00-13:00
Looking forward to meeting you in Berlin!
Early 2018

First, we have released GUI editing alongside text editing for JSON and YAML files on the GIN Web interface.
Secondly, we have added the ability to add DOI files (i.e. datacite.yml) directly to repositories, without copy and paste. This nicely integrates with the GUI YAML editing as our DOI files are YAML files! We hope this does make the process of registering a doi more straightforward.
Thirdly we have added term suggestion to the data search. Give it a try by typing something with at least three letters!
We have also added a cookie warning (which ironically sets an additional cookie btw.)! Be warned!
Finally, we are very happy to announce that that after Nature, now also PLOS has included GIN in their list of recommended repositories. Another major step forward for us!
With that, we hope you had a great start in 2018 and wish you, belatedly, all the best for it (and of course many papers for which you host the data on GIN)!!
December

GIN now has an inbuilt JSON viewer/editor based on the amazing JSON Editor Online from Jos the Jong. Just click on a .json file and you'll see it coming :-).
But what if we had not provided a link to a JSON file above? How could you have found one? That's where the real news is.
We are happy to announce that we have finally launched the GIN Data Search Facilities, which enable you, among other things, to search for files matching some criterion. For example, you could have searched for all JSON files with a query search and the query: "Path:*.json". But that's just the beginning. Basically, you can search for everything now, including the content of files. Try it out with a search for Spike Sorting or Electric Fish. The data search can be found under the explore tab and is equipped with the ability for wildcard and fuzzy searching. Complex search queries are also supported out of the Box!
The technology behind the search we call gindex. Gindex is based on modern search technologies like elasticsearch and it basically analyses repositories for content types it knows (Text, XML, PDF, JSON, odML, NEV to name a few ). From there, it builds a representation that is good for (full-text) searching. You can find more details and documentation here.
As a special Christmas gift, we have also added a Commit search which might make your data organization a bit more flexible :-).
With that, we wish everyone a Merry Christmas and the proverbial happy New Year!!
November 2017
Busy autumn for GIN!
Many things are moving forward for GIN this month and there is more news coming soon, but let's move to today's topic.
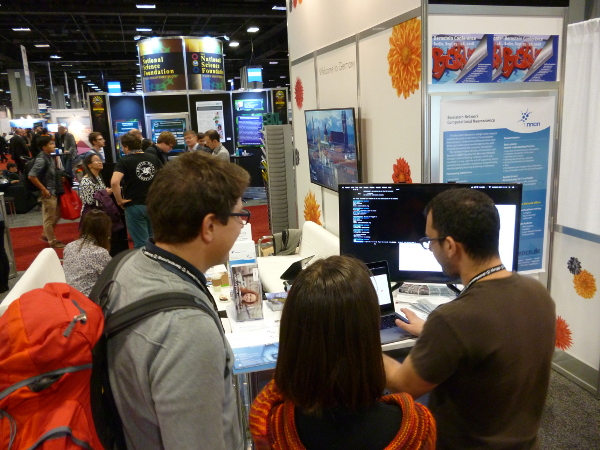

Specifically for this event, we introduced our brand new ARM docker image! We did not want to bring a big server machine to the conference (well you know they are not exactly hand luggage) and therefore, we looked for a somewhat smaller and more mobile solution! Guess what? GIN performs nicely on a Raspberry Pi which, besides being a little bit of a toy deployment for us, is actually nicely illustrating GINs low resource footprint!
There have also been a lot of changes on the server side and quite a bit of our day to day housekeeping has been streamlined. We have moved our weblogs analysis from Webalizer to live-logging with GoAccess. We also migrated our monitoring (well we duplicated it, at least for the time being...) from Nagios to a combination of telegraf, influxDB and Grafana. So at "mission control", which is where we keep an eye (and a notification service) on the GIN-Server metrics, everything is new and shiny!
Finally, Louis has joined the GIN team and will soon help us improving our service with new tutorials and better usage instructions. Welcome to the team!
October 2017
October has been a busy month for GIN and there have been a number of updates:

GIN-Web has been updated with small help messages. Also, we have changed the name of the DOI request file to datacite.yml to increase transparency. We took this opportunity to redesign the DOI UI a bit. For modern browsers code highlighting has been moved to web workers and finally, a lot of tutorial text and imagery has been added.
This brings us to the next point. Kateryna has joined the GIN team and will be working on Tutorials, user instructions, community building, and UI improvements. Welcome to the team!
Finally, we are excited to announce that G-Node is now listed as a recommended data store on the Scientific Data repositories list and on Springer Nature. We hope this will further our mission - to bring easier data sharing and reproducibility to scientist from neuroscience, and the general scientific community!