Readme
This repository contains data resources accompanying the article:
C. Ledig, A. Schuh, R. Guerrero, R. Heckemann, D. Rueckert
"Structural brain imaging in Alzheimer's disease and mild cognitive impairment: biomarker analysis and shared morphometry database", Scientific Reports, 2018.
In this article, we employed a recently validated method (MALPEM) for robust cross-sectional and longitudinal segmentation of MR brain images from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) cohort. Specifically, we segmented 5074 MR brain images into 138 anatomical regions and extracted time-point specific structural volumes and volume change during follow-up intervals of 12 or 24 months.
The work was done in the BioMedIA group at Imperial College London, UK.
Additional experiments and known limitations are collected below.
Citation
Please cite the article as:
@article{Ledig2018,
title={Structural brain imaging in Alzheimer's disease and mild cognitive impairment: biomarker analysis and shared morphometry database},
author={Ledig, Christian and Schuh, Andreas and Guerrero, Ricardo and Heckemann, Rolf A. and Rueckert, Daniel},
journal={Scientific Reports},
doi={10.1038/s41598-018-29295-9},
year={2018},
publisher={Nature Publishing Group}
}
The dataset can be cited as:
@misc{Ledig2018_dataset,
title={Dataset - Structural brain imaging in Alzheimer's disease and mild cognitive impairment: biomarker analysis and shared morphometry database},
author={Ledig, Christian and Schuh, Andreas and Guerrero, Ricardo and Heckemann, Rolf A. and Rueckert, Daniel},
doi={10.12751/g-node.aa605a},
year={2018},
publisher={G-Node},
howpublished= {\url{http://doi.org/10.12751/g-node.aa605a}}
}
File Description
Processed Images from the ADNI cohort
Features
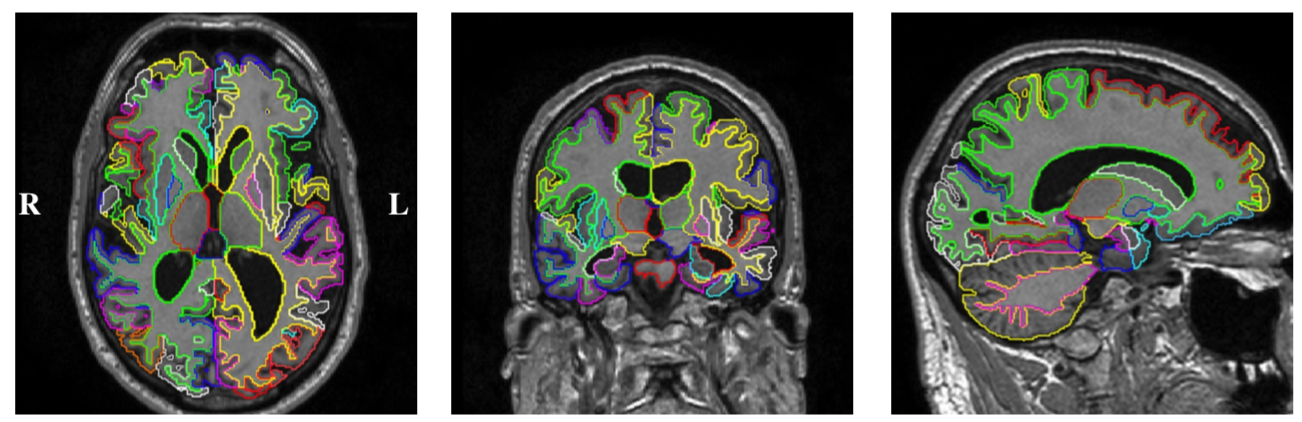
Segmentations / Brain masks
License

The data in this repository is distributed under the terms of the Creative Commons Attribution-NonCommercial 4.0 International Public License. See the accompanying license file for details. The license does not allow usage of this data for commercial applications. This restriction is derived from the license of the Neuromorphometrics atlases (CC BY-NC).
Methodology
Multi-Atlas Label Propagation with Expecation-Maximisation based refinement (MALPEM) including pincram brain extraction is publicly available on github: [MALPEM]
We are working on releasing the actual source code of MALPEM within [MIRTK]
Additional experiments and known limitations
- [Experiment/Limitation - Correction of nuisance factors] -
In the corresponding article, the normalization coefficients for the correction for nuisance factors were calculated on all analyzed healthy control subjects. This allows consistent feature correction across all experiments and simplifies statistical group comparison. We have confirmed that the simpler correction approach we took does not cause any substantial quantitative differences in our experiments.
Specifically, we confirmed that quantitative differences in the calculated statistics are negligible when calculating normalization coefficients based on either the healthy control group or the stable MCI group. Both normalization approaches yield a substantial improvement over uncorrected features.
References
Framework and cross-sectional segmentation [paper]
C. Ledig, R. A. Heckemann, A. Hammers, J. C. Lopez, V. F. J. Newcombe, A. Makropoulos,
J. Loetjoenen, D. Menon and D. Rueckert,
"Robust whole-brain segmentation: Application to traumatic brain injury",
Medical Image Analysis, 21(1), pp. 40-58, 2015.
Longitudinal segmentation [paper]
C. Ledig, W. Shi, A. Makropoulos, J. Koikkalainen, R. A. Heckemann, A. Hammers, J. Lötjönen, and D. Rueckert,
"Consistent and robust 4D whole-brain segmentation: application to traumatic brain injury", Proceedings of ISBI
2014, pp. 673-676, 2014
Brain extraction [paper]
R. Heckemann, C. Ledig, K. R. Gray, P. Aljabar, D. Rueckert, J. V. Hajnal, and A. Hammers,
"Brain extraction using label propagation and group agreement: pincram",
PLoS ONE, 10(7), pp. e0129211, 2015.
Acknowledgements
Data used in preparation of this article were obtained from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database [link]. As such, the investigators within the ADNI contributed to the design and implementation of ADNI and/or provided data but did not participate in analysis or writing of this report. A complete listing of ADNI investigators can be found at: [link]

